|
Medical Imaging Interaction Toolkit
2016.11.0
Medical Imaging Interaction Toolkit
|
|
Medical Imaging Interaction Toolkit
2016.11.0
Medical Imaging Interaction Toolkit
|
The dicom import does not cover all hardware manufacturers but only Siemens dicom images. MITK-DI is also capable of reading the nrrd format, which is documented elsewhere [1, 2]. These files can be created by combining the raw image data with a corresponding textual header file. The file extension should be changed from *.nrrd to *.dwi or from *.nhdr to *.hdwi respectively in order to let MITK-DI recognize the diffusion related header information provided in the files.
In case your dicom images are readable by MITK-DI, select one or more input dicom folders and click import. Each input folder must only contain DICOM-images that can be combined into one vector-valued 3D output volume. Different patients must be loaded from different input-folders. The folders must not contain other acquisitions (e.g. T1,T2,localizer).
In case many imports are performed at once, it is recommended to set the the optional output folder argument. This prevents the images from being kept in memory.
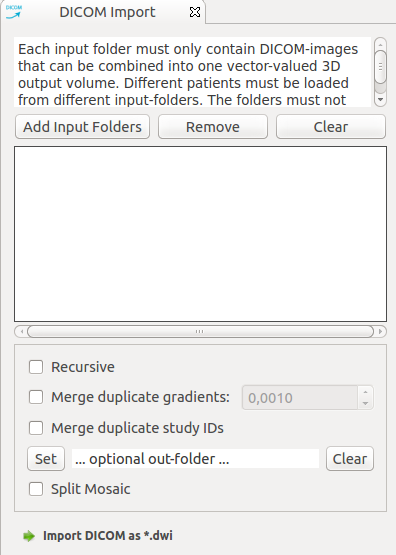
The option "Average duplicate gradients" accumulates the information that was acquired with multiple repetitions for one gradient. Vectors do not have to be precisely equal in order to be merged, if a "blur radius" > 0 is configured.
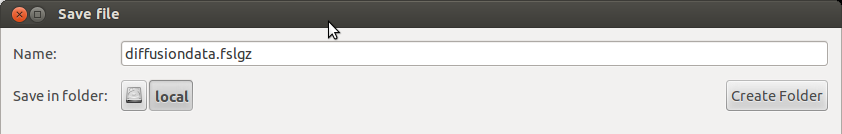
FSL diffusion data can be imported with MITK Diffusion. FSL diffusion datasets consist of 3 files: a nifty file (filename.nii.gz or filename.nii), a bvecs file (filename.bvecs), which is a text file containing the gradient vectors, and a bvals file (filename.bvecs), containing the b-values. Due to the system that selects suitable file readers, MITK will not recognize these files as diffusion datasets. In order to make MITK recognize it as diffusion, the extension must be changed from .nii.gz to .fslgz (so the new name is filename.fslgz) or from filename.nii to filename.fsl. The bvecs and bvals files have to be renamed as well(to filename.fsl.bvecs/filenames.fsl.bvecs or to filename.fslgz.bvecs/filename.fslgz.bvals).
MITK can also save diffusion weighted images in FSL format. To do this the extension of the new file should be changed to .fsl or .fslgz upon saving the file.